Noel Lowndes

Prof Noel Lowndes
Established Professor of Biochemistry
Director of the CCB
SFI Future Frontiers & EMBO Member
noel.lowndes@universityofgalway.ie
Research interests
- Role of phosphatidylinositol 3-kinase like kinases (PIKKs) in genome stability
- Kinesins & 53BP1-dependent DNA double strand break repair
- Zinc finger protein roles in DNA double strand break signalling & repair
- Regulation of cytokinetic abscission by ATR
Research overview
Pathways required to maintain genome stability are potently tumour suppressive, while genome instability is a hallmark of cancer. Our research aims to elucidate novel mechanisms controlling genome stability. While our focus is on fundamental cell biology, future translational studies may ultimately have clinical impact. We utilise genome editing, proteomics, in vitro biochemistry and microscopy to identify and dissect novel genome stability factors. Our principle models are established transformed and immortalised human cell lines (e.g. HCT293, HeLa, U2OS, hTERT-RPE1 cells). In addition, we use primary animal fibroblasts (e.g. mouse, bat).
Keywords
ATM, ATR, abscission, checkpoints, DNA repair, kinesins, 53BP1
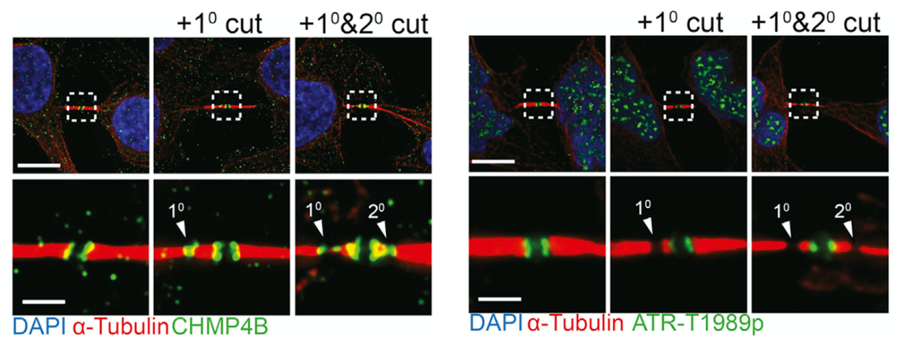
Selected Figures
Key Research techniques
- Genome editing
- Protein biochemistry & proteomics
- DNA damage & cell cycle assays
- Fluorescence & expansion microscopy
Lab Members
- Dr Ilze Skujina (Post-Doc)
- Dr Sabrinna Crowley (Post Doc)
- Peng Wu (PhD)
- Anna-Marie Meaney (PhD)
- Zaid Abu Diak (PhD)
Mentoring/Hosting

Dr Janna Luessing
Senior Researcher
SFI-IRC Pathway Programme
janna.luessing@universityofgalway.ie
.
Selected publications
- The nuclear kinesin KIF18B promotes 53BP1-mediated DNA double-strand break repair
- A role for the p53 tumour suppressor in regulating the balance between homologous recombination and non-homologous end joining
- ATR activates the S-M checkpoint during unperturbed growth to ensure sufficient replication prior to mitotic onset
- Multiple facets of the DNA damage response contribute to the radioresistance of mouse mesenchymal stromal cell lines
- Budding yeast Rad9 is an ATP-dependent Rad53 activating machine
Most Recent Publications
Quick Links
Get in Touch!
noel.lowndes@universityofgalway.ie